|
5/2/2023 0 Comments Imagej thresholdingThe latter option is also recommended if one needs to execute the macro multiple times. The macro editor also provides a convenient Run button to start the macro. Furthermore, if you want to edit the default parameters that are displayed in the graphical interface or edit any other aspect of the macros, got to Plugins > Macros > Edit. and choose the macro file you want to execute. To execute a macro in ImageJ/Fiji, simply click Plugins > Macros > Run. A thorough introduction to the use and development of macros can be found here. Macros are simple scripts that execute a series of ImageJ-commands/functions sequentially and may feature a graphical interface for the user to enter parameters. Again, the user may review the automatic detection and separation of the nuclei, and change any erroneous detections before the automatic analysis. To separate clumps of nuclei, the macro applies Watershed Separation. It can be used to check the number of nuclei expressing a certain protein, and to estimate the corresponding expression level for individual nuclei. The macro subsequently measures the intensity of those nuclei in a second fluorescence channel.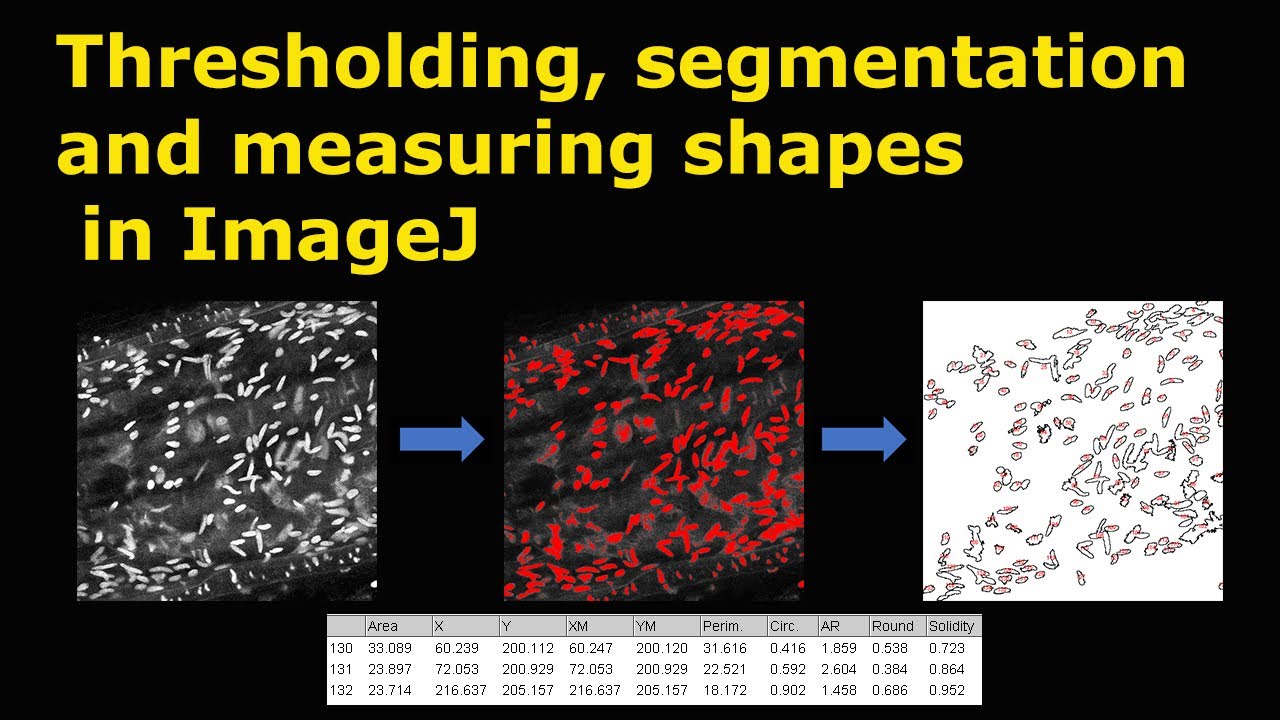 
NucleiFluorescenceIntensity.ijm is a more specialized version of the first macro, and focuses on detecting cell nuclei in one fluorescence channel (tested with DAPI). Furthermore, a second series of images can be supplied to automatically analyze the intensity values in these images, based on the segmented regions. This repository contains two different macros (click on filename to download):įluorescenceIntensityAnalysis.ijm is a general macro to threshold (fluorescent) objects in images and automatically analyze the area fraction covered by those objects in a series of images, or measure the area of individual objects.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
